(Preprint) Metagenomic evidence for the coexistence of SARS and H1N1 in patients from 2007-2012 flu seasons
Qi Liu ¹ ² ⁴, Zhenglin Du 杜政霖 ² ³ ⁴, Sihui Zhu ² ³ ⁴, Wenming Zhao 赵文明 ² ³ ⁴, Hua Chen 陈华 ¹ ² ⁴, Yongbiao Xue 薛勇彪 ² ³ ⁴
¹ CAS Key Laboratory for Genomic and Precision Medicine, Beijing Institute of Genomics, Chinese Academy of Sciences, Beijing 100101, China. 中国科学院 北京基因组研究所 精准基因组医学重点实验室
² China National Center for Bioinformation, Beijing 100101, China. 国家生物信息中心
³ National Genomics Data Center, Beijing Institute of Genomics, Chinese Academy of 10 Sciences, Beijing 100101, China. 北京基因组研究所 国家基因组科学数据中心
⁴ University of Chinese Academy of Sciences, Beijing 100049, China. 中国科学院大学
ChinaXiv, 2021-08-11
Abstract
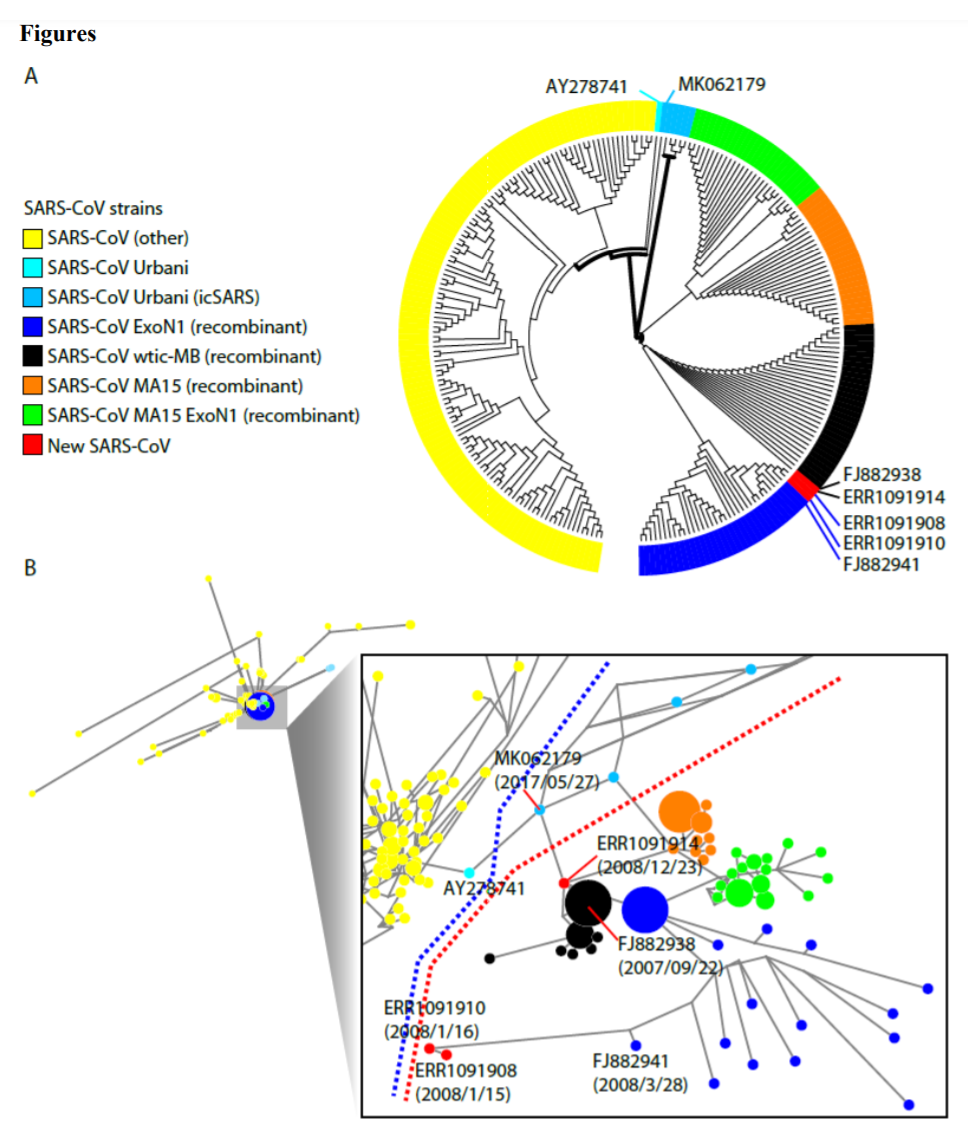
By re-analzying public metagenomic data from 101 patients infected with influenza A virus during the 2007-2012 H1N1 flu seasons in France, we identified 22 samples with SARS-CoV sequences. In 3 of them, the SARS genome sequences could be fully assembled out of each. These sequences are highly similar (99.99% and 99.7%) to the artificially constructed recombinant 5 SARS-CoV (SARSr-CoV) strains generated by the J. Craig Venter Institute in USA.
Moreover, samples from different flu seasons have different SARS-CoV strains, and the divergence between these strains cannot be explained by natural evolution. Our study also shows that retrospective studies using public metagenomic data from past major epidemic outbreaks serves as a genomic strategy for the research of origins or spread of infectious diseases.
A 4096-element 3D-integrated Si-SiN optical phased array for high-power coherent LiDAR
Han Wang, Weimin Xie, Xin Yan, Jiaqi Li, Youxi Lu, Ping Jiang, Feng Li, Kai Jin, Xu Yang, Jiali Jiang, Keran Deng, Weishuai Chen, Jing Luo, Li Jin, Junbo Feng, Kai Wei
Opto-Electronic Technology
2026-03-20
High-speed and large-capacity visible light communication for 6G: advances and perspectives
Nan Chi, Zhilan Lu, Fujie Li, Haoyu Zhang, Yunkai Wang, Xinyi Liu, Zhiwu Chen, Zhe Feng, Zhuoran Hu, Zhixue He, Ziwei Li, Chao Shen, Junwen Zhang
Opto-Electronic Technology
2026-03-20
Holotomography-driven learning unlocks in-silico staining of single cells in flow cytometry by avoiding fluorescence co-registration
Daniele Pirone, Giusy Giugliano, Michela Schiavo, Annalaura Montella, Martina Mugnano, Vincenza Cerbone, Maddalena Raia, Giulia Scalia Ivana Kurelac, Diego Luis Medina, Lisa Miccio Mario Capasso, Achille Iolascon, Pasquale Memmolo, Pietro Ferraro
Opto-Electronic Science
2026-02-25